4.2 Gene Variants
What is a Gene Variant and how do Variants Occur?
Variant vs. Mutation
What is a genomic variant?
All humans have near-identical DNA sequences across the estimated six billion-letter code for their genome. Slight differences exist between individuals, making each of us unique. These differences, called genomic variants, occur at specific locations within the DNA. DNA is read like a code. Recall this code is made up of four types of chemical building blocks – adenine, thymine, cytosine and guanine, abbreviated with the letters A, T, C and G. A genomic variant occurs in a location within the DNA where that code differs among people. For example, in Person One below, the location shows a “C” base. But in the exact location in Person Two, it is a “T.”
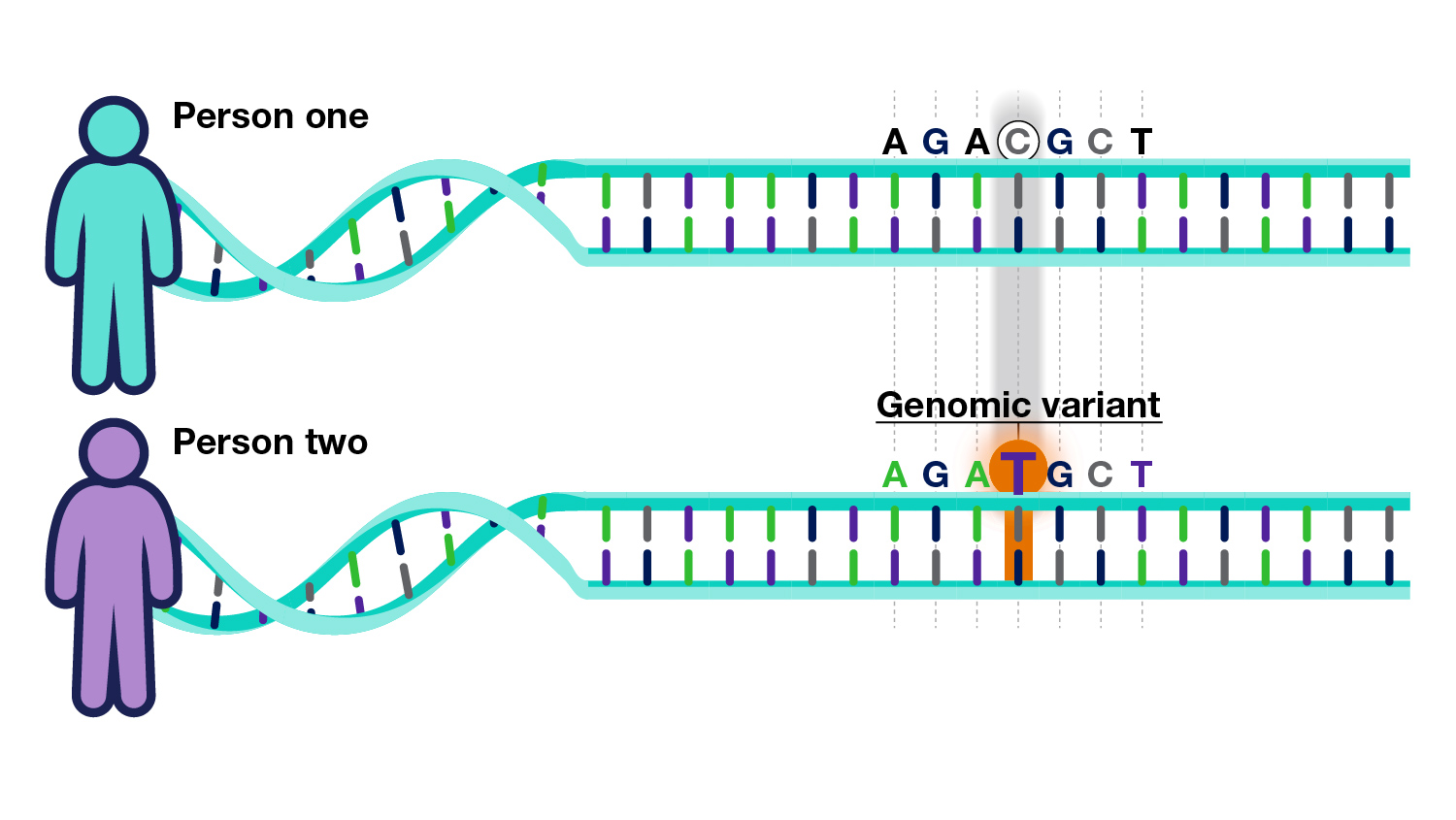
An individual’s genome has roughly 4 to 5 million such genomic variants. These variants may be unique to that individual or occur in others. Some variants increase the risk of developing diseases, while others may reduce such risk; others do not affect disease risk. In other words,
The question is: How do these genomic variants influence the risk for specific diseases?
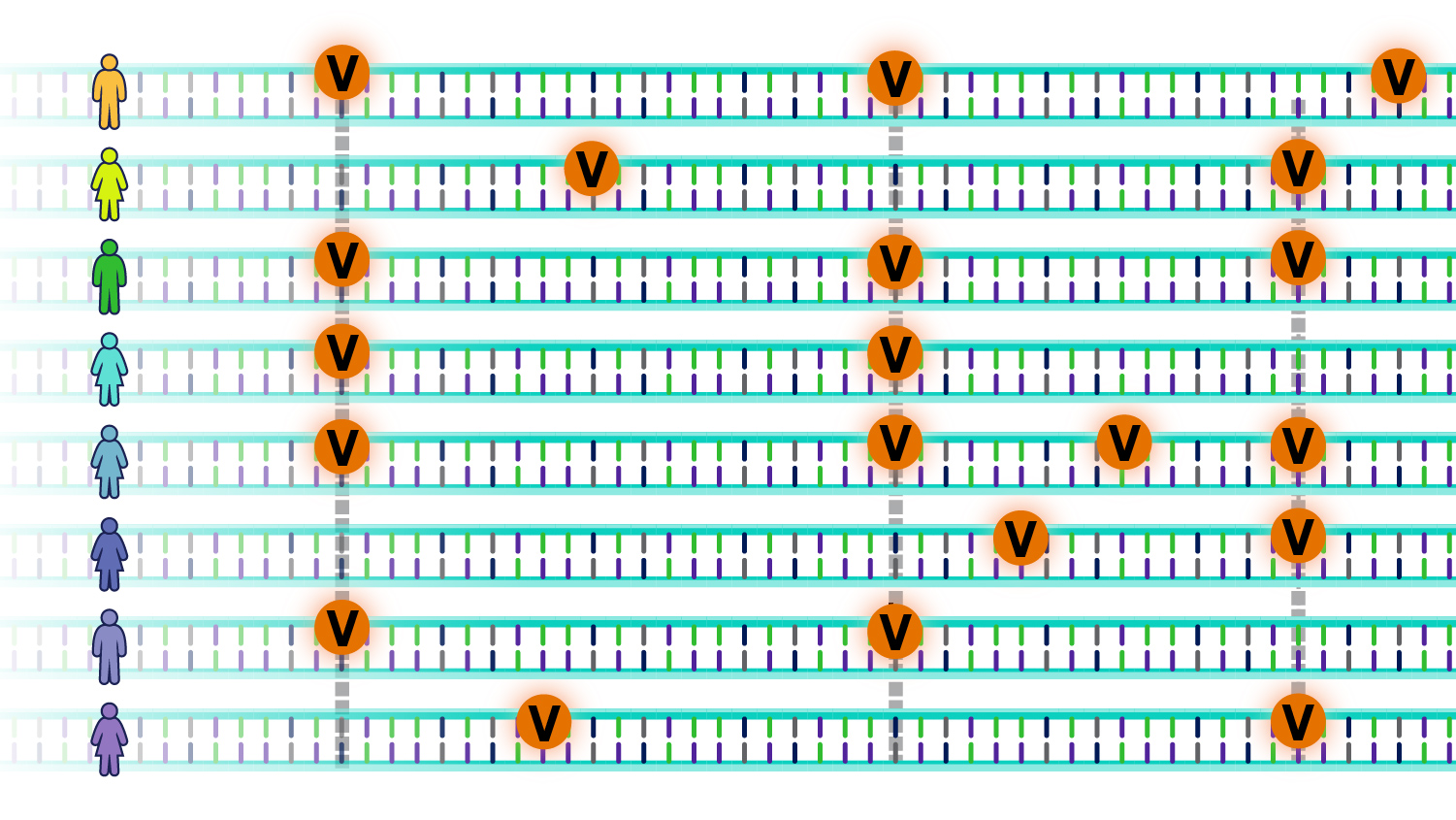
Morphological Variants
Morphological variants cause changes in the visible form of the organism as they give rise to altered forms of a trait, e.g., change in size, shape (normal wing vs. curly wing in fruit flies), colour, number, etc.

Lethal Variants
A lethal variant causes the premature death of an organism. For example, in Drosophila (fruit flies), lethal variants can result in death during the embryonic, larval, or pupal stages. Lethal variants are usually recessive, so both gene copies must be lost for premature death (homozygous lethal alleles will not be viable). Heterozygotes, which have one lethal allele and one wild-type allele, are typically viable. In the example shown in the figure below regarding yellow coat colour in mice, the lethal allele is recessive because it causes death only in homozygotes. Unlike its effect on survival, the effect of the allele on colour is dominant. A single copy of the allele in heterozygotes produces a yellow colour in mice. These examples illustrate the point that the type of dominance depends on the aspect of the phenotype examined.
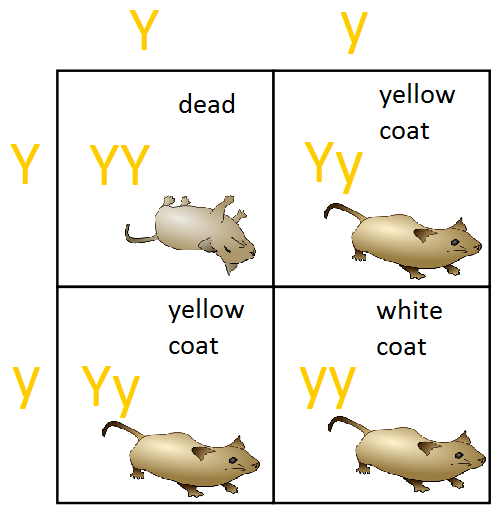

Conditional Variants
Conditional variations rely on phenotype = genotype + environment + interaction. Organisms with this variant express an altered phenotype, but only under specific environmental conditions. Under restrictive conditions, they express the altered phenotype; under permissive conditions, they show a wild-type phenotype. One example of a conditional variant is the temperature-sensitive pigmentation of Siamese cats. Siamese cats have a temperature-sensitive fur colour; their fur appears unpigmented (light-coloured) when grown in a warm environment. The hair appears pigmented (dark) when grown at a cooler temperature. This is seen at the peripheral regions of the feet, snout, and ears. This is because, in warm temperatures, the enzyme needed for melanin pigment synthesis becomes nonfunctional. However, in cooler temperatures, the enzyme required for melanin synthesis is functional, and the deposition of melanin makes the fur look dark.
Do all gene variants affect health and development?
A very small percentage of all variants actually have a positive effect (gain-of-function variant). These variants lead to new versions of proteins that help an individual better adapt to environmental changes. For example, a beneficial variant could result in a protein that protects an individual and future generations from a new strain of bacteria.
Gene variants can be inherited from a parent or occur during a person’s lifetime:
- Inherited (or hereditary) variants are passed from parent to child and are present throughout a person’s life in virtually every cell in the body. These variants are also called germline variants because they are present in the parent’s egg or sperm cells, which are also called germ cells. When an egg and a sperm cell unite, the resulting fertilized egg cell contains DNA from both parents. Any variants in that DNA will be present in the child’s cells that grow from the fertilized egg.
- Non-inherited variants occur at some time during a person’s life and are present only in certain cells, not in every cell in the body. Because non-inherited variants typically occur in somatic cells (cells other than sperm and egg cells), they are often called somatic variants. These variants cannot be passed to the next generation. Non-inherited variants can be caused by environmental factors such as sun ultraviolet radiation or an error when DNA copies itself during cell division.
Some genetic changes are described as new (de novo) variants; these variants are recognized in a child but not in either parent. The variant sometimes occurs in a parent’s egg or sperm cell but is absent in other cells. In other cases, the variant occurs in the fertilized egg shortly after the egg and sperm cells unite. (It is often impossible to tell exactly when a de novo variant happened.) As the fertilized egg divides, each resulting cell in the growing embryo will have the variant. De novo variants are one explanation for genetic disorders in which an affected child has a variant in every cell in the body. Still, the parents do not, and there is no family history of the disorder.
Variants acquired during development can lead to mosaicism, in which a set of cells in the body has a different genetic makeup than others. In mosaicism, the genetic change is not present in a parent’s egg or sperm cells or the fertilized egg but happens later, anytime from embryonic development through adulthood. As cells grow and divide, cells that arise from the cell with the altered gene will have the variant, while other cells will not. It is called somatic mosaicism when a proportion of somatic cells have a gene variant and others do not. Depending on the variant and how many cells are affected, somatic mosaicism may or may not cause health problems. When a proportion of egg or sperm cells have a variant and others do not, it is called germline mosaicism. In this situation, an unaffected parent can pass a genetic condition to their child.
Most variants do not lead to disease development, and those that do are uncommon in the general population. Some variants occur often enough in the population to be considered common genetic variation. Several variants are responsible for differences between people, such as eye colour, hair colour, and blood type. Although many of these common variations in the DNA have no adverse effects on a person’s health, some may influence the risk of developing certain disorders.
Silent Changes
After mutagen treatment, most base pair changes (especially substitutions) have no noticeable effect on the phenotype. Often, this is because the change occurs in the DNA sequence of a non-coding region of the DNA, such as in the intergenic areas (between genes) or within an intron where the sequence does not code for protein and is not essential for proper mRNA splicing. Also, even if the change affects the coding region, it may not alter the amino acid sequence (recall that the genetic code is degenerate; for example, GCT, GCC, GCA, and GCG all encode alanine) so a change in a single nucleotide may result in a codon that produces the same amino acid. This is referred to as a silent variant. The base substitution may also change an amino acid, but this does not quantitatively or qualitatively alter the product’s function so that no phenotypic change would occur.
Environment and Genetic Redundancy
In some situations, a variant can cause a complete loss of function of a gene yet not produce a change in the phenotype, even when the mutant allele is homozygous. The lack of a visible phenotypic change can be due to environmental effects: losing that gene product may not be apparent in that specific environment but might be in another.
Alternatively, the lack of a phenotype might be attributed to genetic redundancy. That is, the mutant gene’s lost function is compensated by another gene at another locus, encoding a similarly functioning product. Thus, the loss of one gene is compensated by the presence of another. The concept of genetic redundancy is an essential consideration in genetic screens. A gene whose function can be compensated for by another gene cannot be easily identified in a genetic screen for loss-of-function variants.
How can gene variants affect health and development?
To function correctly, each cell depends on thousands of proteins to do their jobs in the right places at the right times. A variant can cause a protein normally expressed by a gene to malfunction or to not be produced at all. When a variant alters a protein that plays a critical role in the body, it can disrupt normal development or cause a health condition. A condition caused by variants in one or more genes is called a genetic disorder.
It is important to emphasize that genes do not cause disease—genetic disorders are caused by variants that alter or eliminate a gene’s function. For example, when people say someone has “the cystic fibrosis gene,” they usually refer to a version of the CFTR gene that contains a variant that causes the disease. All people, including those without cystic fibrosis, have a version of the CFTR gene.
Essential Genes and Lethal Alleles
In some cases, gene variants are so severe that they prevent an embryo from surviving until birth (lethal variants). Sometimes, it is required to reach a particular developmental stage before the phenotype resulting from a variant can be seen or scored. For example, flower colour can only be scored in plants that are mature enough to make flowers, and eye colour can only be scored in flies that have developed to the adult stage. However, some organisms with variants may not develop sufficiently to reach a stage that can be scored for a particular phenotype. Variants in essential genes create recessive lethal alleles that arrest or derail the development of an individual at an immature (embryonic, larval, or pupal) stage. Therefore, this variant type may go unnoticed in a typical variant screen because they are absent from the screened progeny. Furthermore, the progeny of a monohybrid cross involving an embryonic lethal recessive allele may all be of a single phenotypic class, giving a phenotypic ratio of 1:0 (which is the same as 3:0). In this case, the variant may not be detected. Nevertheless, studying recessive lethal variants (those in essential genes) has elucidated many important biochemical pathways.
What kinds of gene variants are possible?
The human genome’s most common polymorphisms (or genetic differences) are single base-pair differences. Scientists call these differences SNPs for single-nucleotide polymorphisms. When two different haploid genomes are compared, SNPs occur, on average, about every 1,000 bases, other types of polymorphisms—for example, differences in copy number, insertions, deletions, duplications, and rearrangements—also occur, but much less frequently (NIH & BSCS, 2007).
Note: The images in the following section still use the term “mutation.”
The DNA sequence of a gene can be altered in a number of ways. Gene variants can have varying effects on health, depending on where they occur and whether they alter the function of essential proteins. Variant types include the following:
Substitution
This variant type replaces one DNA building block (nucleotide) with another. Substitution variants can be further classified by their effect on protein production from the altered gene.
- Missense:A missense variant is a type of substitution in which the nucleotide change replaces one protein building block (amino acid) with another in the protein made from the gene. The amino acid change may alter the function of the protein.
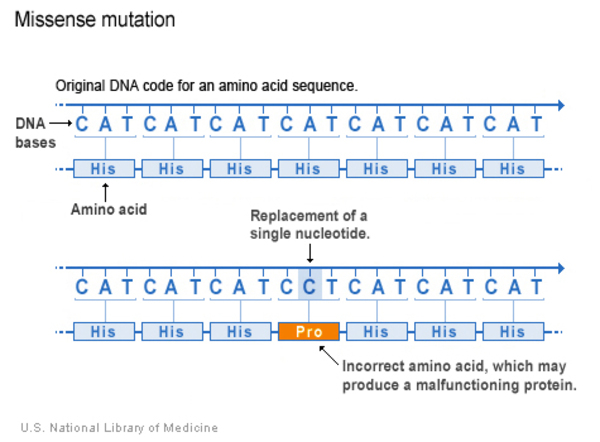
- Nonsense: A nonsense variant is another type of substitution. However, instead of causing a change in one amino acid, the altered DNA sequence results in a stop signal that prematurely signals the cell to stop building a protein. This type of variant results in a shortened protein that may malfunction, be nonfunctional or get broken down.
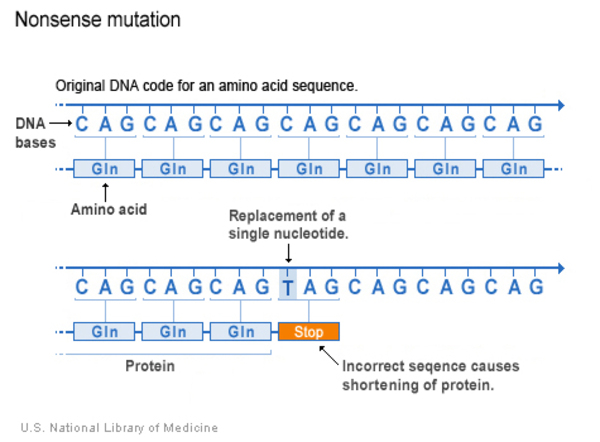
Insertion
An insertion changes the DNA sequence by adding one or more nucleotides to the gene. As a result, the protein made from the gene may not function properly.
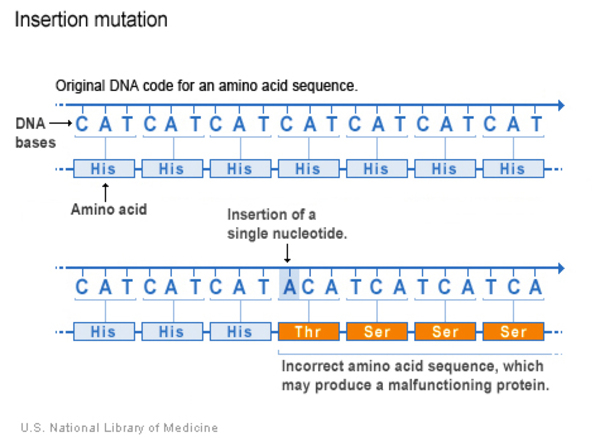
Deletion
A deletion changes the DNA sequence by removing at least one nucleotide in a gene. Small deletions remove one or a few nucleotides within a gene, while larger deletions can remove an entire gene or several neighbouring genes. The deleted DNA may alter the function of the affected protein or proteins.
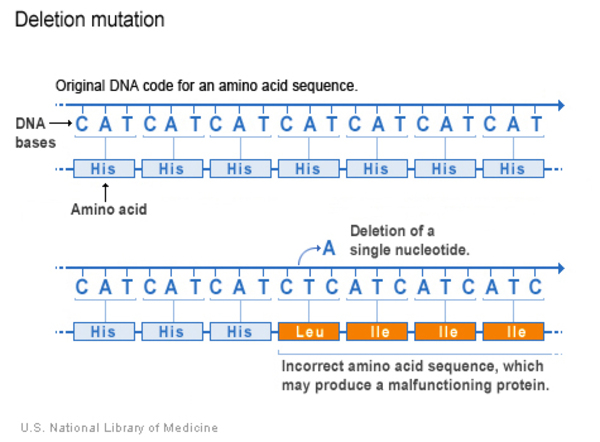
Deletion-Insertion
This variant occurs when a deletion and insertion happen at the same time in the same location in the gene. In a deletion-insertion variant, at least one nucleotide is removed and at least one nucleotide is inserted. However, the change must be complex enough to differ from a simple substitution. The resulting protein may not function properly. A deletion-insertion (delins) variant may also be called an insertion-deletion (indel) variant.
Duplication
A duplication occurs when a stretch of one or more nucleotides in a gene is copied and repeated next to the original DNA sequence. This type of variant may alter the function of the protein made from the gene.
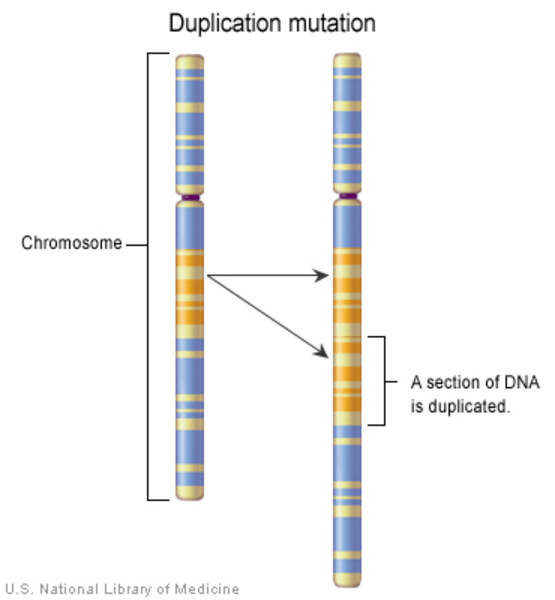
Inversion
An inversion changes more than one nucleotide in a gene by replacing the original sequence with the same sequence in reverse order. We will discuss this further in the subsequent chapter on chromosomal disorders.
Frameshift
A reading frame consists of groups of three nucleotides that each code for one amino acid.
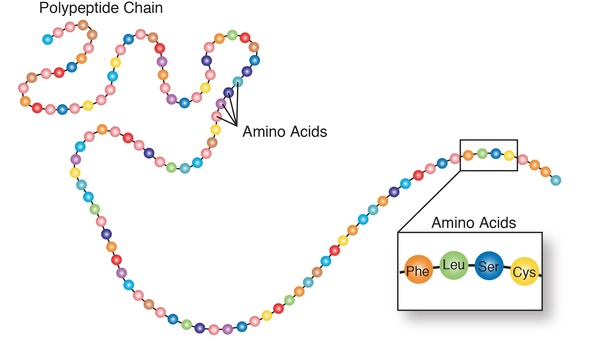
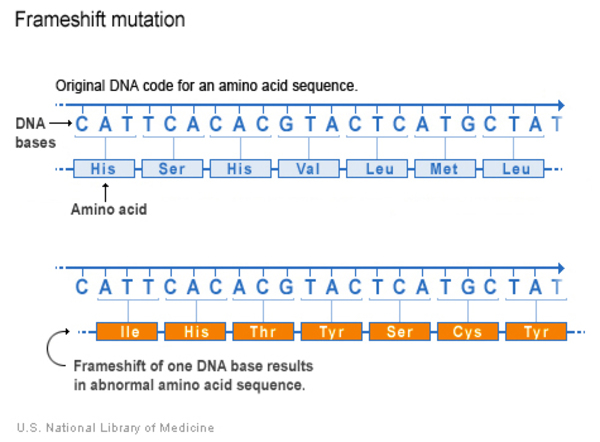
A frameshift variant occurs when there is an addition or loss of nucleotides that shifts the grouping and changes the code for all downstream amino acids. The resulting protein is usually nonfunctional. Insertions, deletions, and duplications can all be frameshift variants.
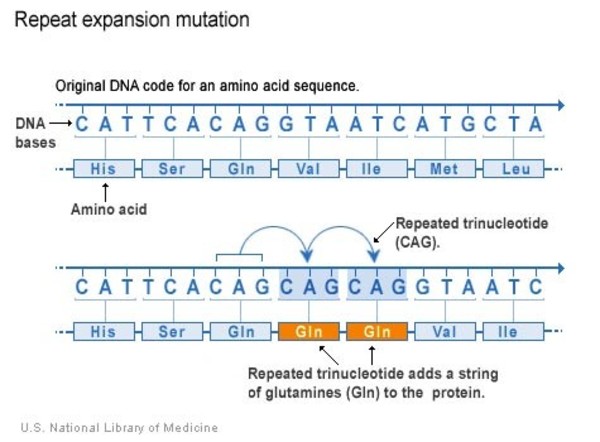
Repeat expansion
Some regions of DNA contain short sequences of nucleotides that are repeated a number of times in a row. For example, a trinucleotide repeat is made up of sequences of three nucleotides, and a tetranucleotide repeat is made up of sequences of four nucleotides. A repeat expansion is a variant that increases the number of times that the short DNA sequence is repeated. This type of variant can cause the resulting protein to malfunction.
Can a change in the number of genes affect health and development?
People have two copies of most genes, one copy inherited from each parent. In some cases, however, the number of copies varies—meaning that a person can have one, three, or more copies of particular genes. Less commonly, both copies of a gene may be missing. These genetic differences are known as copy number variations (CNV).
Copy number variation results from insertions, deletions, and duplications of large segments of DNA that are at least one thousand nucleotides (also called one kilobase or 1kb) in length. These segments are often big enough to include whole genes. Variations in gene copy number can influence the activity of genes and the functioning of proteins made from them, which may affect body processes.
Copy number variation accounts for a significant amount of genetic difference between people. More than 10 percent of the human genome appears to contain differences in gene copy number. While much of this variation does not affect health or development, some differences influence a person’s risk of disease, particularly some types of cancer, or response to certain drugs.
Attribution & References
- 6.7 Classification of Mutants In Introduction to Genetics by Natasha Ramroop Singh, Thompson Rivers University, CC BY-NC-SA 4.0
- Do all gene variants affect health and development? In Help Me Understand Genetics by MedlinePlus, National Library of Medicine, Public Domain with attribution
- 7.5 Mutations Without Detectable Phenotypes In Introduction to Genetics by Natasha Ramroop Singh, Thompson Rivers University, CC BY-NC-SA 4.0
- How can gene variants affect health and development? In Help Me Understand Genetics by MedlinePlus, National Library of Medicine, Public Domain with attribution
- What kinds of gene variants are possible? In Help Me Understand Genetics by MedlinePlus, National Library of Medicine, Public Domain with attribution
- What is a genomic variant? Courtesy: National Human Genome Research Institute, Public domain with attribution
- Can a change in the number of genes affect health and development? In Help Me Understand Genetics by MedlinePlus, National Library of Medicine, Public Domain with attribution
References
National Institutes of Health (US) (NIH), & Biological Sciences Curriculum Study (BSCS). (2007). Understanding human genetic variation. NIH Curriculum Supplement Series. National Institutes of Health (US). https://www.ncbi.nlm.nih.gov/books/NBK20363/

