3.7 Epigenetics in Practice
The Impact on Health and Clinical Practice
Certain diseases can change your epigenetics. In addition, some epigenetic changes can make you more likely to develop certain diseases, such as cancer.
Infections (CDC, n.d.)
Germs can change your epigenetics to weaken your immune system. This helps the germ survive.
EXAMPLE: Mycobacterium tuberculosis causes tuberculosis. Infections with these germs can cause epigenetic changes in some of your immune cells that result in turning off the IL-12B gene. Turning off the IL-12B gene weakens your immune system and improves the survival of Mycobacterium tuberculosis (Chandran et al., 2015).
Cancer (CDC, n.d.)
Certain gene variants make you more likely to develop cancer. Likewise, some epigenetic changes increase your cancer risk. Most cancers display different epigenetic patterns than those of healthy cells, and these patterns can be important for the development of the disease. For example, some genes produce proteins that stop cell growth and division. If expression is suppressed in these genes when it shouldn’t be, this could cause cells to divide uncontrollably, leading to cancer development. Additionally, having a mutation in the BRCA1 gene that prevents it from working properly makes you more likely to get breast and other cancers. Similarly, increased DNA methylation that results in decreased BRCA1 gene expression raises your risk for breast and other cancers (Tang et al., 2016). While cancer cells have increased DNA methylation at certain genes, overall DNA methylation levels are lower in cancer cells compared with normal cells. These differences can help us to understand, manage and treat cancers more effectively. However, epigenetics alone cannot diagnose cancer. Cancers would need to be confirmed with further screening tests. An example of the use of epigenetics in cancer diagnostics is the methylation profile of tumours can help to grade the current stage of a cancer or to track the disease as it progresses over time. Further, many drugs have been discovered that target specific epigenetic modifications, either by removing them or preventing their removal – depending on the desired effect. These drugs are mostly used to treat different types of cancer, but there is ongoing work to establish their effectiveness for other conditions.
EXAMPLE: Colorectal cancers have abnormal DNA methylation near certain genes, which affects expression of these genes. Some commercial colorectal cancer screening tests (for example, Cologuard®) use stool samples to look for this abnormal DNA methylation. It is important to know that if the result of one of these tests is positive or abnormal, further screening is required with a colonoscopy (Chan & Liang, 2022).
Epigenomic studies spotlight new level of cancer evolution (NHS, 2023)
Two studies cast a light on the role of epigenomics in oncology. A multi-omic level of cancer evolution has been characterized by researchers from the Institute of Cancer Research. In a pair of Nature publications, they highlighted the importance of epigenomic changes to a cancer’s resilience, and how genome-only testing methods may be missing important cancer markers. Every cancer is different, yet even within one mass, its cells are not identical: the cells can differ by which genes they express and their susceptibility to treatment. Both studies examined the evolution of colorectal cancers and tried to understand why cells within a cancer differ in terms of what drives cancer growth and cancer cell survival.
Epigenomic changes are common
In the first of the two studies, researchers looked at 30 patients with bowel cancer. They examined each cancer’s genome and its epigenome. Using this multi-omic approach, they found that changes to the epigenome were common around a cancer’s driver genes. Cancer driver genes are genes that are normally involved in a cell’s division and growth but gene variants alter their normal functioning. Additionally, the epigenomic changes were found to be passed on to the next generation of cells; they were present in cancer cells that had survival advantages over other cells.
Tumours persist due to variation across its cells
Cells within a cancer mass can often be different from each other. The second study took a deep dive into the differences found within tumours. The researchers did this by sampling many different sections of individual tumours to understand how and why the cells of a single cancer can be so diverse. They found that, while there is variation in the expression of genes within tumours, in most cases this doesn’t alter the ability of cells to survive or grow. However, if the cancer’s environment changes (such as during anti-cancer treatment), then this variation becomes important for its future survival as it may mean that some of its cells survive while others do not. Ultimately, it’s the cancer’s surviving cells that pass on their resistance to treatment to their daughter cells, meaning the cancer, in some form, persists.
“For years our understanding of cancer has focused on gene variants which permanently change the DNA code,” said Institute of Cancer Research director and lead author Professor Trevor Graham, “But our research has shown that the way the DNA folds up can change which genes are read without altering the DNA code, and this can be very important in determining how cancers behave.”
How is this being used in practice?
Knowledge around cancer epigenomics and how it fuels cancer growth is already being used in cancer detection. For example, the Galleri multi-cancer test, uses epigenomic markers to detect non-symptomatic cancers. Yet the epigenome is not widely being used in precision cancer treatment. If it was, Professor Graham noted that, “we could, potentially, much more accurately predict which treatments will work best for a particular person’s cancer.” Tools are available that could make personalized cancer treatment a reality, such as long-read sequencing approaches, like nanopore, which allow real-time ‘reading’ of DNA to understand the genome, and parts of the epigenome too.
To learn more about the latest advances in epigenetics and its real-world impact on healthcare, visit the dedicated epigenomics topic on the NHS Genomics Education Programme’s blog.
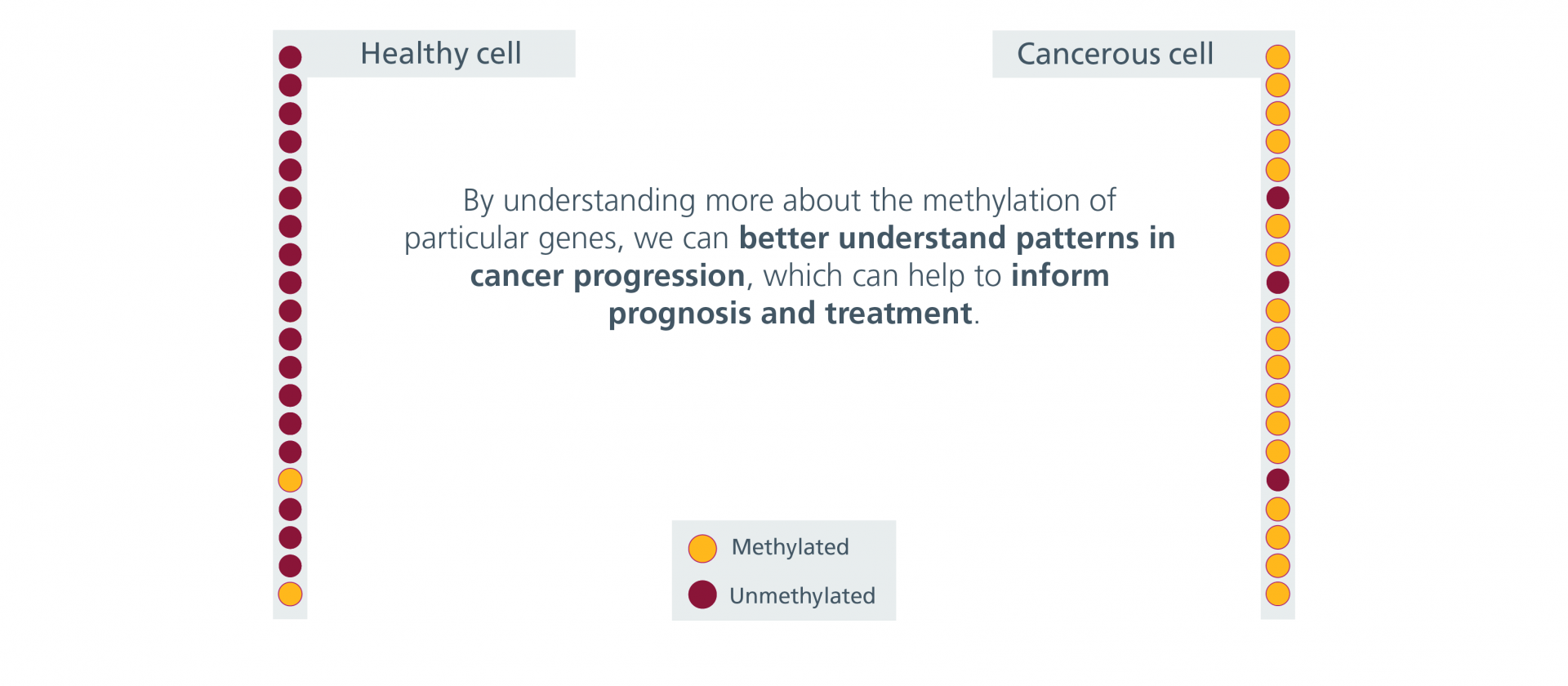
Check out the gallery below to see how epigenetic patterns can change between healthy cells and cancer cells.
Epigenetic Patterns in Health and Cancerous Cells – Gallery
Epigenetic Patterns in Health and Cancerous Cells – Gallery (text version)
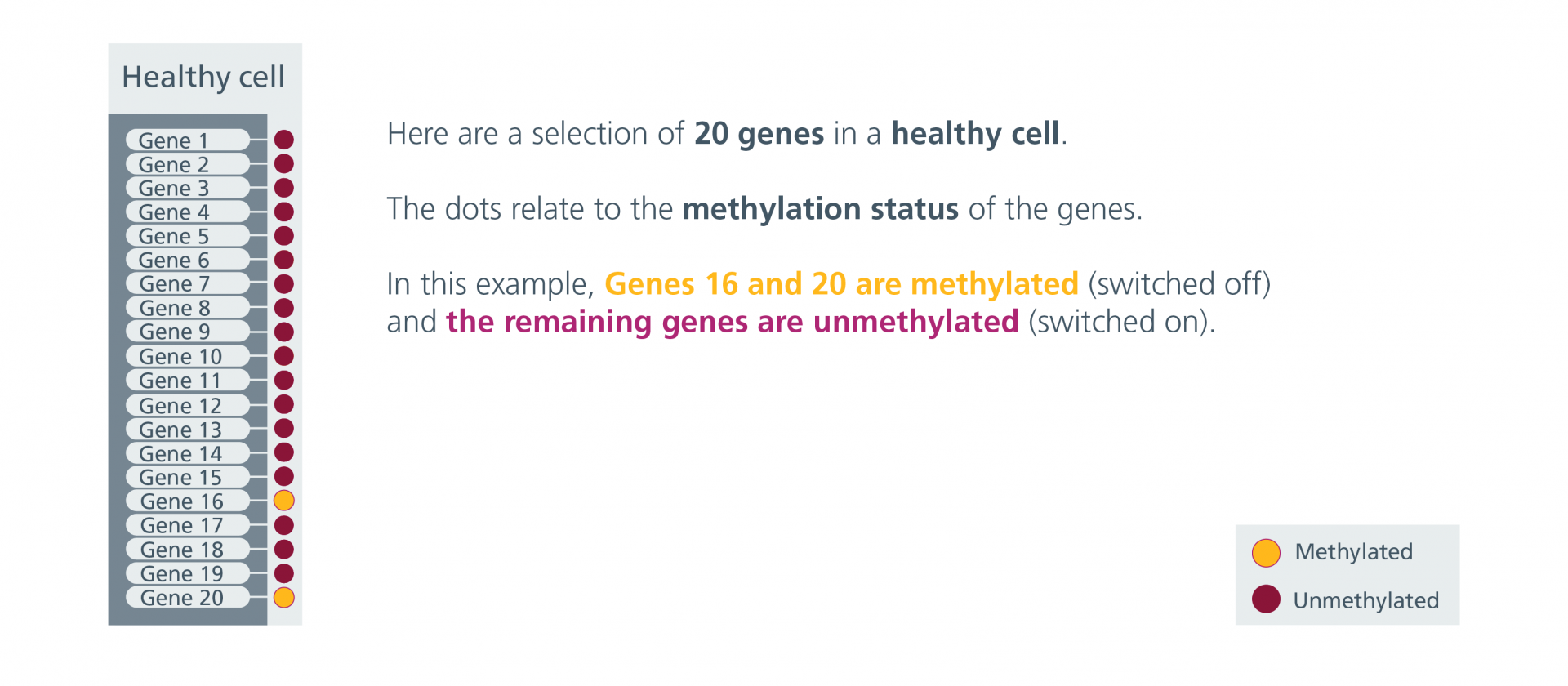
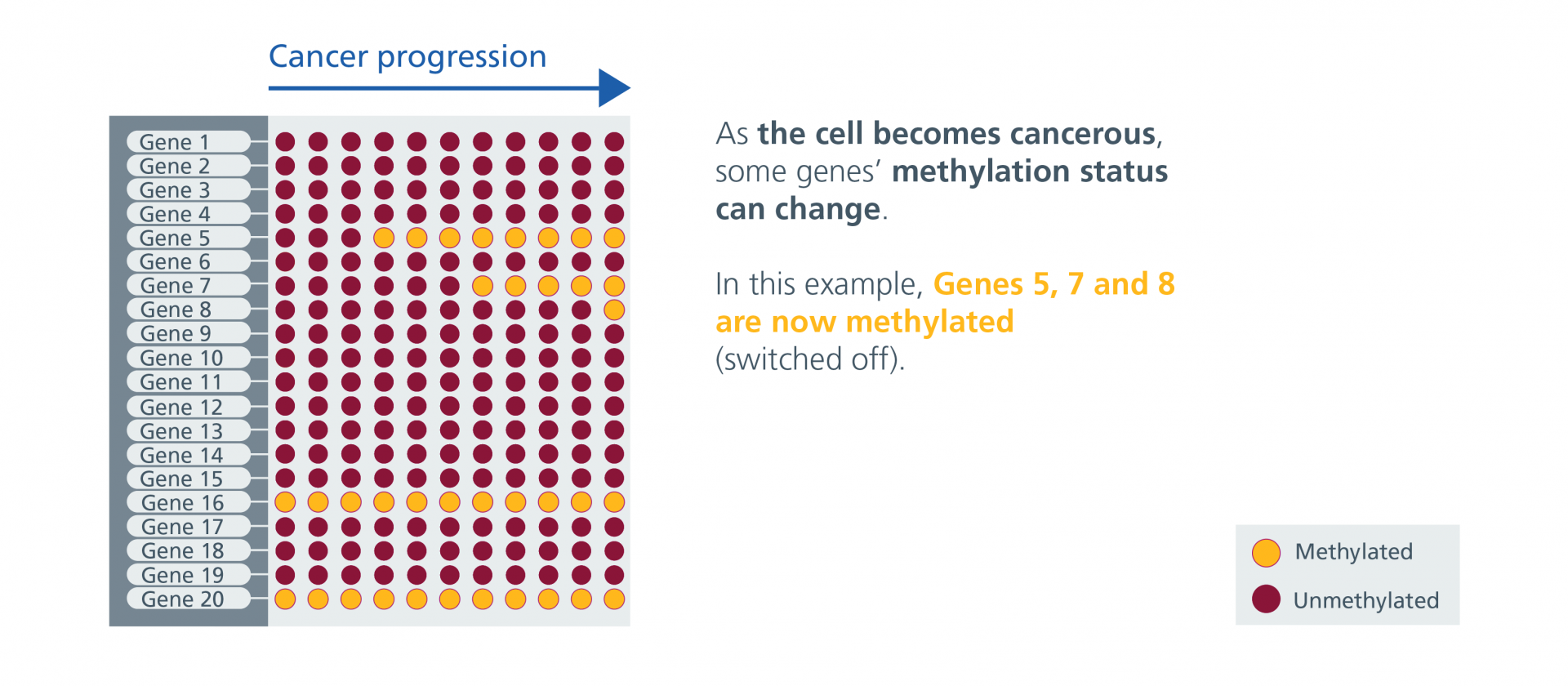
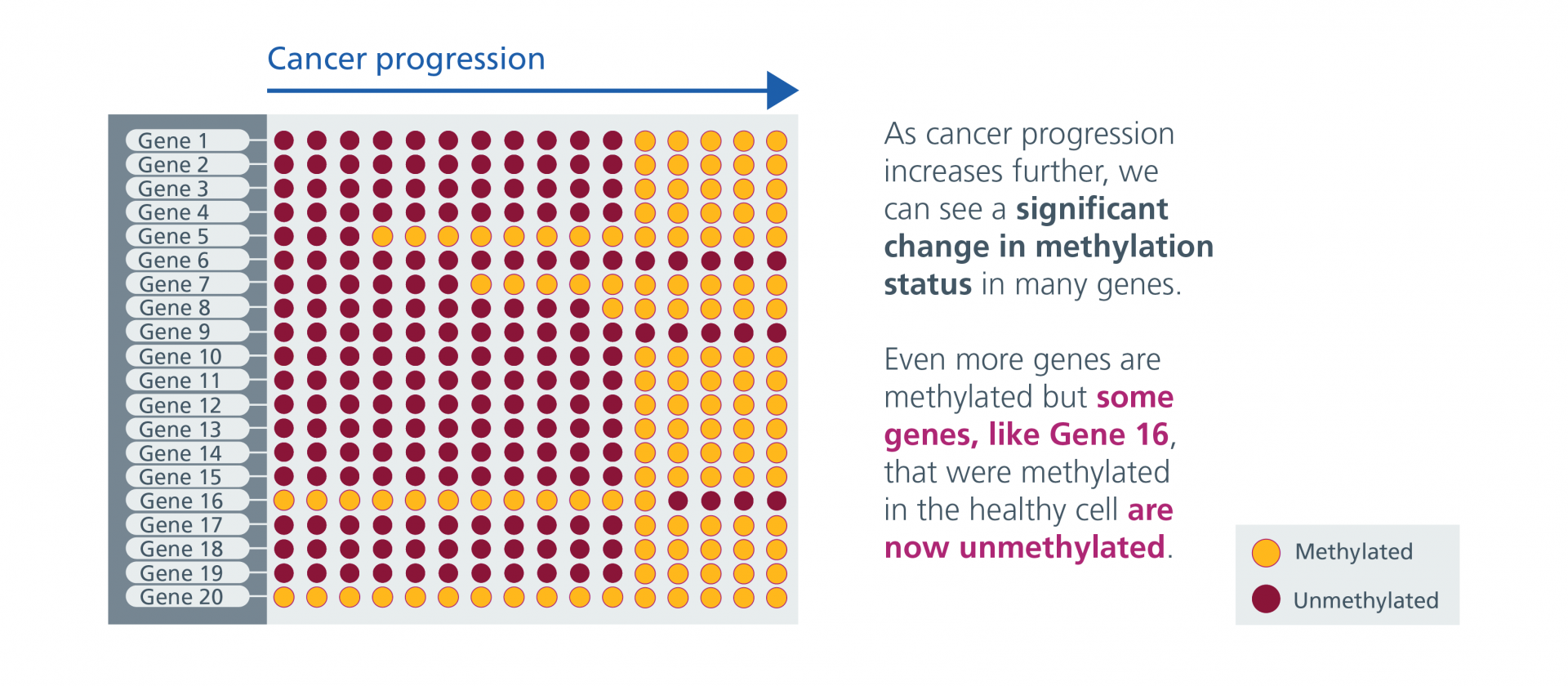

Source: Genomics Education Programme, CC BY-NC 4.0
Imprinting disorders
Individually a rare disease is rare, but collectively they are common. Many people with rare diseases often go through a diagnostic odyssey with many tests that do not necessarily lead to an explanation or treatment for their disease. It is thought that the majority of rare diseases have a genetic cause. Finding this cause can lead to a diagnosis and possibly treatment options. Increasingly, more people with undiagnosed rare diseases are undergoing genomic testing which is providing a diagnosis for some. Not everyone who has a genomic test will receive a diagnosis initially, but those without a diagnosis may receive one as more is discovered about the function of the genome.
A variety of factors can cause rare diseases. Many occur because of alterations to the DNA sequence itself, but some can be caused by epigenetic modifications: in other words, changes in gene expression. Genomic imprinting is one such modification that affects the regulation of certain genes, but how can it cause disease?
As we have learned, epigenetic changes happen because our gene expression is altered by various mechanisms. This suppression or amplification of gene expression is very common, and indeed is an important part of normal, healthy gene regulation. However, if this regulation is disrupted, it can lead to certain genes being over or under expressed, or silenced completely.
Although many of our genes are affected by epigenetic modifications, only a very small percentage are known to undergo genomic imprinting. These genes are mostly found in two clusters – one on the short arm of chromosome 11 and the other on the long arm of chromosome 15.
How genomic imprinting works
Genomic imprinting refers to the process by which certain genes are ‘branded’ with the parent of origin. When gametes (sperm and eggs) are made, epigenetic markers that were inherited from our parents or accumulated in life are removed, but in genes that undergo genomic imprinting, new markers are added that identify the gene as coming from either the mother or the father.
These new markers change gene expression, resulting in the imprinted copy of the gene being turned off and the other copy being turned on. For example, if the allele inherited from the father is imprinted, it is silenced and only the allele from the mother is expressed; if the allele from the mother is imprinted, then only the allele from the father is expressed.
Uniparental disomy
Genomic imprinting can cause disease when there are errors in gamete production, or during early embryonic development. One common complication is uniparental disomy (UPD), which is when a person inherits two copies of a chromosome from one parent, and none from the other.
When the chromosome in question does not contain imprinted regions, UPD may have no deleterious effects, but when chromosomes 11 or 15 are involved, the situation can be more complex. Because some genes in these regions are inactivated by maternal or paternal imprinting, an individual who inherits two copies from one parent can lack an active copy of some essential genes – even though they are present in the genome.
Which conditions can it cause?
The first conditions that were identified as being caused by imprinting were Angelman Syndrome and Prader-Willi Syndrome. Both conditions are caused by missing genetic information on chromosome 15 – one of the chromosomes where genomic imprinting is common.
Angelman syndrome occurs when the UBE3A gene is not working properly or is missing entirely. The paternal copy of the gene is imprinted, or silenced, meaning that only the copy inherited from the person’s mother is active. Problems with or deletion of the maternal copy of the gene, or the section of chromosome 15 where it resides, can lead to Angelman syndrome.
In Prader-Willi syndrome (PWS), the same region of chromosome 15 is involved. In this case, though, the condition is linked to problems with the paternal copy of the chromosome rather than the maternal copy. Certain genes in the region, such as SNRPN, are imprinted, with the maternal copy being silenced and the paternal copy being active. Problems with the paternal copies of these genes can lead to PWS.
Watch the video on Prader-Willi syndrome (bottom of the page) from the NHS to learn about the features of PWS.
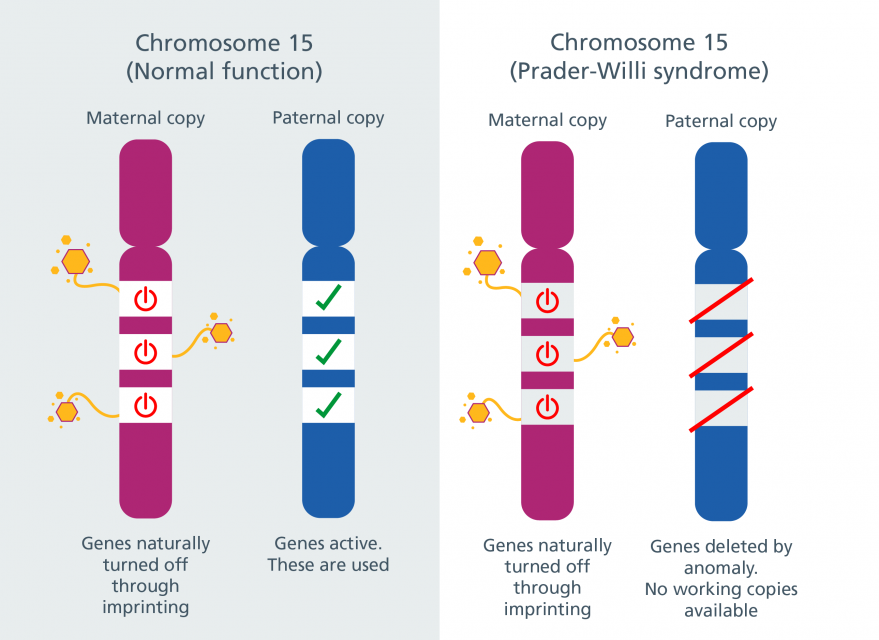
Most cases of Angelman syndrome and PWS are caused by deletions, although a significant minority are due to UPD.
Another condition linked to genomic imprinting is Beckwith-Wiedemann syndrome (BWS). It is caused by changes to imprinted genes in the chromosome 11 cluster, with a number of genes being implicated thus far, including H19 and IGF2. It is associated with abnormal growth and an increased chance of childhood cancers, especially in the liver and kidneys.
Scientists hope that continued research into the genetic pathways involved in BWS will help to improve treatment methods and outcomes for those with the condition.
Implications for Policy: Social and Environmental Epigenetics
DNA methylation serves as a biological record of early social or environmental conditions, offering valuable insights for policy and interventions aimed at improving the social and structural determinants of health. Policy implications include (Schmidt, 2019):
- Guiding Policy Directions: Epigenetic research highlights the long-term health impact of early-life environments, similar to how evidence of second-hand smoke risks led to public smoking bans. Raising awareness about these biological effects could support child-focused interventions and policies that regulate environmental (e.g. air pollution) and social exposures (e.g. food security). For example, stricter regulations on industrial emissions or agricultural chemicals may be justified by findings demonstrating their long-term health impacts through epigenetic mechanisms.
- Biomarkers for Early Screening: Epigenetics may offer biomarkers for identifying children at risk of health conditions, like fetal alcohol spectrum disorder (FASD), allowing for screening and early interventions that maximize health outcomes as well as targeted health campaigns.
- Evaluating Intervention Effectiveness: Epigenetic markers may contribute to the assessment of the biological impact of interventions long before physical health outcomes manifest, enabling quicker and more cost-effective policy evaluations.
- Shaping Global Environmental and Health Strategies: As epigenetic research works toward finding evidence of how environmental factors affect health across generations, it has the potential to shape global agreements and policies. Collaborative efforts could focus on reducing environmental health disparities, addressing climate change, and prioritizing sustainable practices that protect vulnerable populations.
Quiz – Units 1-3
See the Blackboard course shell for the syllabus which provides details about the quiz content and due dates.
Attribution & References
Except where otherwise noted, this content is adapted from:
- What is Epigenetics by Genomics Education Programme, CC BY-NC
- Epigenomic studies spotlight new level of cancer evolution by Genomics Education Programme, CC BY-NC
- What is genomic imprinting? by Genomics Education Programme, CC BY-NC
- Epigenetics, Health, and Disease by CDC, Public Domain with attribution, disclaimer
- Implications for Policy by Andrea Gretchev, CC BY-NC
Adaptation notes: Use of CDC material does not imply endorsement by CDC. Material is otherwise available on the CDC website free of charge.
References
Chan, S. C. H., & Liang, J. Q. (2022). Advances in tests for colorectal cancer screening and diagnosis. Expert review of molecular diagnostics, 22(4), 449–460. https://doi.org/10.1080/14737159.2022.2065197
Chandran, A., Antony, C., Jose, L., Mundayoor, S., Natarajan, K., & Kumar, R. A. (2015). Mycobacterium tuberculosis infection induces HDAC1-mediated suppression of IL-12B gene expression in macrophages. Frontiers in Cellular and Infection Microbiology, 5, 90. https://doi.org/10.3389/fcimb.2015.00090
Schmidt, K. (2019, November 14). Social epigenetics: How your early life environment gets “under your skin” Policy Horizons Canada. https://horizons.service.canada.ca/en/2019/11/14/social-epigenetics-how-your-early-life-environment-gets-under-your-skin/index.shtml
Tang, Q., Cheng, J., Cao, X., Surowy, H., & Burwinkel, B. (2016). Blood-based DNA methylation as biomarker for breast cancer: a systematic review. Clinical epigenetics, 8, 115. https://doi.org/10.1186/s13148-016-0282-6

